Research Articles
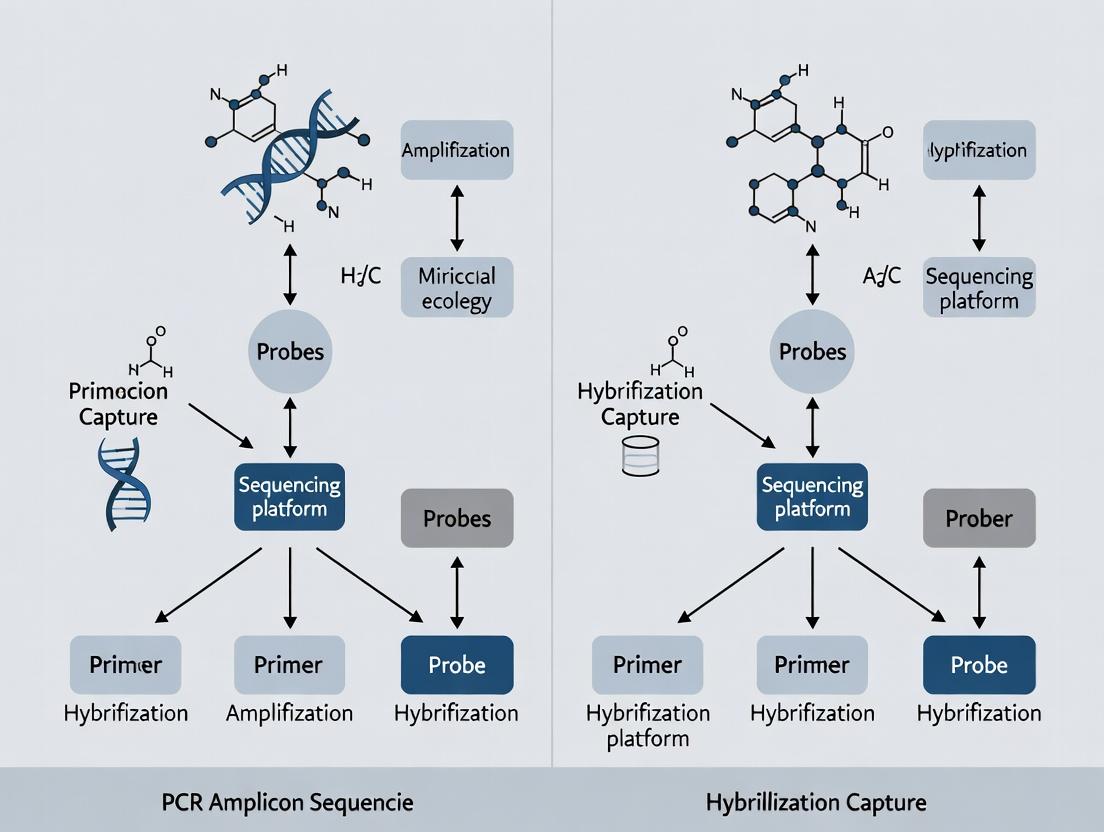
Targeted Sequencing Face-Off: When to Choose PCR Amplicon vs. Hybridization Capture for Precision Variant Detection
This article provides a comprehensive comparison of PCR amplicon sequencing and hybridization capture, the two dominant approaches for targeted next-generation sequencing (NGS) in research and drug development.
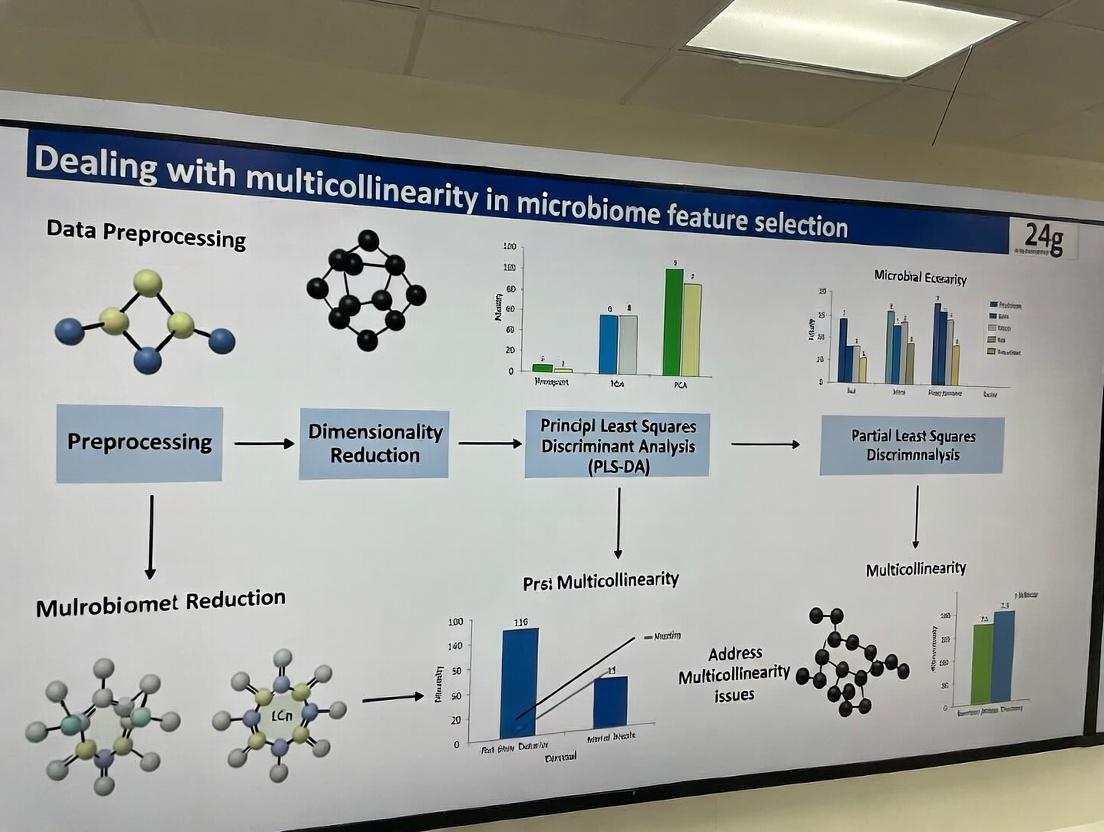
Untangling the Microbial Web: A Practical Guide to Handling Multicollinearity in Microbiome Feature Selection
This article provides a comprehensive guide for researchers and drug development professionals on addressing the pervasive challenge of multicollinearity in microbiome feature selection.
Beyond Spurious Correlations: A Practical Guide to Handling Compositional Bias in Biomedical Data Analysis
This article provides a comprehensive guide for researchers and drug development professionals on identifying and correcting for compositional data bias in correlation analyses.
Beyond Missing Data: A Practical Guide to Advanced Imputation Methods for Sparse Microbiome Analysis in Biomedical Research
Sparse, zero-inflated data is a fundamental challenge in microbiome research, introducing significant bias and hindering downstream statistical and machine learning analyses.
Boosting Microbial Detection: How DOPE-FISH Amplifies Signal Intensity for Researchers
This article provides a comprehensive guide for researchers, scientists, and drug development professionals on the DOPE-FISH (Double Labeling of Oligonucleotide Probes - Fluorescence In Situ Hybridization) technique.
DNA-SIP vs RNA-SIP: A Comparative Guide to Sensitivity, Resolution, and Applications in Microbial Ecology
This article provides a comprehensive comparison of DNA-based Stable Isotope Probing (DNA-SIP) and RNA-based Stable Isotope Probing (RNA-SIP), two pivotal techniques for linking microbial identity to function in complex environments.
Mastering DNA-SIP with 13C: A Comprehensive Protocol for Microbial Functional Identification and Drug Discovery
This article provides a detailed, step-by-step guide to the DNA Stable Isotope Probing (DNA-SIP) protocol utilizing 13C-labeled substrates, tailored for researchers in biomedical science and drug development.
Mastering DNA-SIP Centrifugation: Essential Guide to Density Gradient Challenges, Troubleshooting & Applications
This comprehensive guide addresses the critical challenges and solutions in DNA-Stable Isotope Probing (DNA-SIP) density gradient centrifugation, a key technique for linking microbial identity to function.
Decoding the Living Microbiome: A Comparative Guide to DNA vs. RNA-Based Microbial Community Analysis
This article provides a comprehensive comparative analysis of DNA-based and RNA-based approaches for microbial community composition profiling.
DNA vs RNA 16S Sequencing: Choosing the Right Amplicon Method for Microbiome Research and Drug Discovery
This comprehensive guide explores the critical distinction between DNA-based and RNA-based 16S ribosomal RNA (rRNA) amplicon sequencing in microbiome analysis.